IGV sightseeing tour
In this section we will have a quick tour through several types of genomic sequencing, using data from a single sample.
To run this tutorial you will need the IGV desktop application installed - or work with someone who does.
To get IGV, go to the download page and download and install the appropriate version. (The ones with Java included are simplest, but have a bigger download.)
Start up IGV and make sure you have selected the 'Human (GRCh38/h38)' genome build from the drop-down at the top left:
Now search for a gene by typing it into the search box and pressing <enter> - a good one to start with is FUT2, which is on chromosome 19.
The datasets enclosed are all from a single individual, codenamed healthy volunteer 31 (HV31). If you want to read more about sequencing of that individual, see this paper.
Note
For this practical we've generated files that only cover a subset of genes, in order to keep them small. If you can't see any data at any point, it might be because you're in the wrong part of the genome. To re-orient yourself, try entering one of these genes in the search box:
- FUT1 and FUT2 on chromosome 19
- APOE, also on chromosome 19
- CD14
- UBASH3A
- G6PD
- GNAS
Warning
Please note that all data in this sightseeing tour is included strictly for training purposes - it is not publicly-available data. Please do not share outside this course. Contact me (Gavin Band) if you have any queries about this.
Instructions
We have a set of 7 datasets for you to explore, all from HV31 genome. The first is a paired-end Illumina sequencing data much like what we have been analysing so far today.
For each dataset you'll get the BAM file URL, something like this:
https://www.chg.ox.ac.uk/bioinformatics/training/gms/data/sequence_data_sightseeing_tour/illumina.bam
To load this into IGV, do the following:
- Select 'Load from URL' from the file menu at the top of the screen.
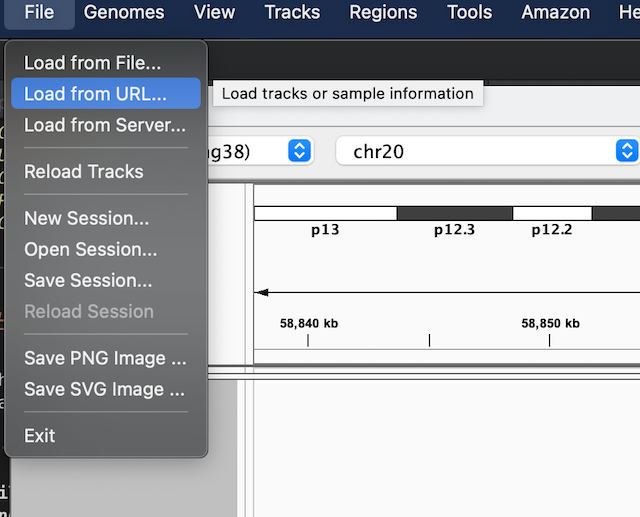
Copy and paste in the dataset URL into the 'File URL' box.
The 'index URL' is just the same as the file url, with '.bai' added. Copy and paste or type it in.
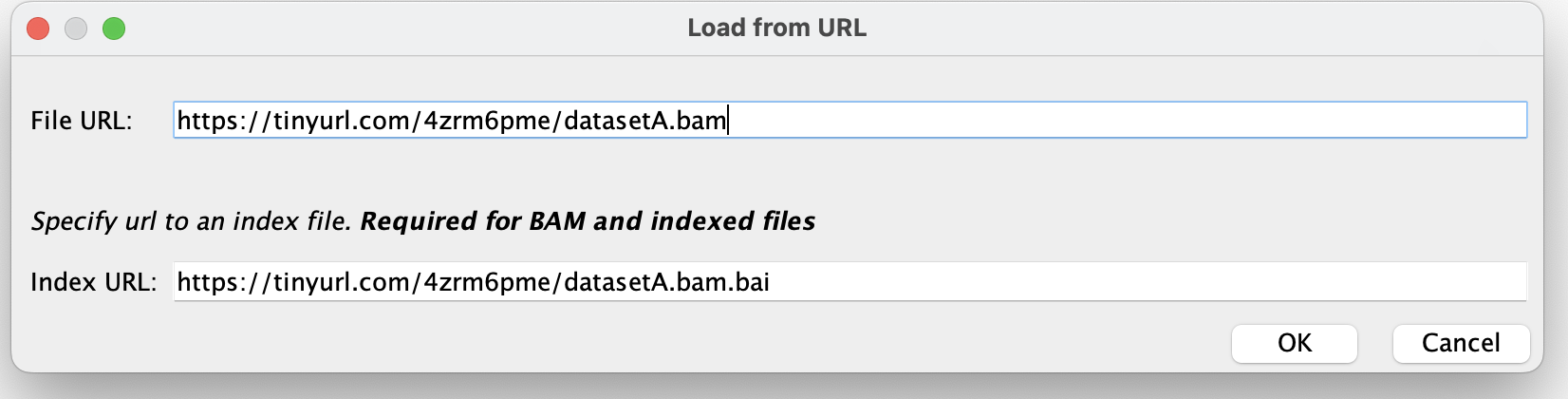
- Press 'OK'
You should see the data loaded into a new track.
The datasets
Ok we are ready! Let's go:
- Load dataset A - Illumina short-read genomic sequencing.
- Load datasets B and C - Nanopore and PacBio long-read genomic sequencing.
- See base modifications in the long-read data.
- Load dataset D - Illumina RNA-seq, from CD14+ monocytes and other cells.
- Load Dataset E - Illumina ATAC-seq.
- Load Dataset F - Illumina sequencing of 10X linked-read libraries.
- New!! Load Dataset G - Pacbio long-read RNA-seq data.
Next steps
When you've had a look it's time to play guess the dataset - good luck!