Getting chromosome lengths
To answer questions like 'how much of the genome is in genes' we need to know what the length of each genome is. Happily, this data is already in the GFF files - though it's a bit awkward to get at. In fact:
- Some of the GFF files have records with
type='chromosome'in, that reflect this data - while others have records with
type='region'. - still others have records with
type='supercontig'. - Some GFF files - like the ones from PlasmoDB - don't have them at all. (Luckily there are files from Ensembl protists which we can use instead.)
Here is some dbplyr code to load these chromosomes.
contigs = (
db
%>% tbl( 'gff' )
%>% filter( type %in% c( 'chromosome', 'region', 'supercontig' ))
%>% select( dataset, ID, type, seqid, source, start, end, attributes )
%>% collect()
)
As a sanity check let's make sure that this really does capture the seqid values for gene records.
We'll use the trusy stopifnot() for this:
table( genes$dataset, genes$seqid %in% contigs$seqid )
TRUE
Acanthochromis_polyacanthus.ASM210954v1.110 24016
Asparagus_officinalis.Aspof.V1.57 24141
Bufo_bufo-GCA_905171765.1-2022_05-genes 20900
Camelus_dromedarius.CamDro2.110.chr 18896
Gallus_gallus.bGalGal1.mat.broiler.GRCg7b.110.chr 16711
Homo_sapiens.GRCh38.110.chr 20046
Mus_musculus.GRCm39.110.chr 21633
Pan_troglodytes.Pan_tro_3.0.110.chr 21879
Plasmodium_falciparum.ASM276v2.57 5358
Plasmodium_knowlesi.ASM635v1.57 5102
Plasmodium_vivax.ASM241v2.57 5389
Good!
This suggests we can compute the genome lengths.
contigs$sequence_length = contigs$end - contigs$start + 1
(
contigs
%>% group_by( dataset, type )
%>% summarise(
number = n(),
length = sum(sequence_length)
)
%>% mutate(
length_in_mb = sprintf( "%.1fMb", length / 1E6 )
)
)
dataset type number length length_in_mb
<chr> <chr> <int> <dbl> <chr>
1 Acanthochromis_polyacanthus.ASM210954v1.110 region 30414 991584656 991.6Mb
2 Asparagus_officinalis.Aspof.V1.57 chromosome 10 1111887942 1111.9Mb
3 Asparagus_officinalis.Aspof.V1.57 supercontig 11782 75650082 75.7Mb
4 Bufo_bufo-GCA_905171765.1-2022_05-genes region 1306 5044744194 5044.7Mb
5 Camelus_dromedarius.CamDro2.110.chr region 37 2052758708 2052.8Mb
6 Gallus_gallus.bGalGal1.mat.broiler.GRCg7b.110.chr region 42 1041139641 1041.1Mb
7 Homo_sapiens.GRCh38.110.chr chromosome 25 3088286401 3088.3Mb
8 Mus_musculus.GRCm39.110.chr chromosome 22 2723431143 2723.4Mb
9 Pan_troglodytes.Pan_tro_3.0.110.chr chromosome 26 2967125077 2967.1Mb
10 Plasmodium_falciparum.ASM276v2.57 chromosome 14 23292622 23.3Mb
11 Plasmodium_knowlesi.ASM635v1.57 chromosome 14 23462187 23.5Mb
12 Plasmodium_vivax.ASM241v2.57 chromosome 14 22621071 22.6Mb
13 Plasmodium_vivax.ASM241v2.57 supercontig 2733 4386630 4.4Mb
Note
You might be wondering what 'region', 'chromosome', and 'supercontig' are - if so, so am I!
Looking up these genomes in NCBI Datasets sheds some light on this. For example, try looking up the Plasmodium vivax assemblyy. You should see something like this:
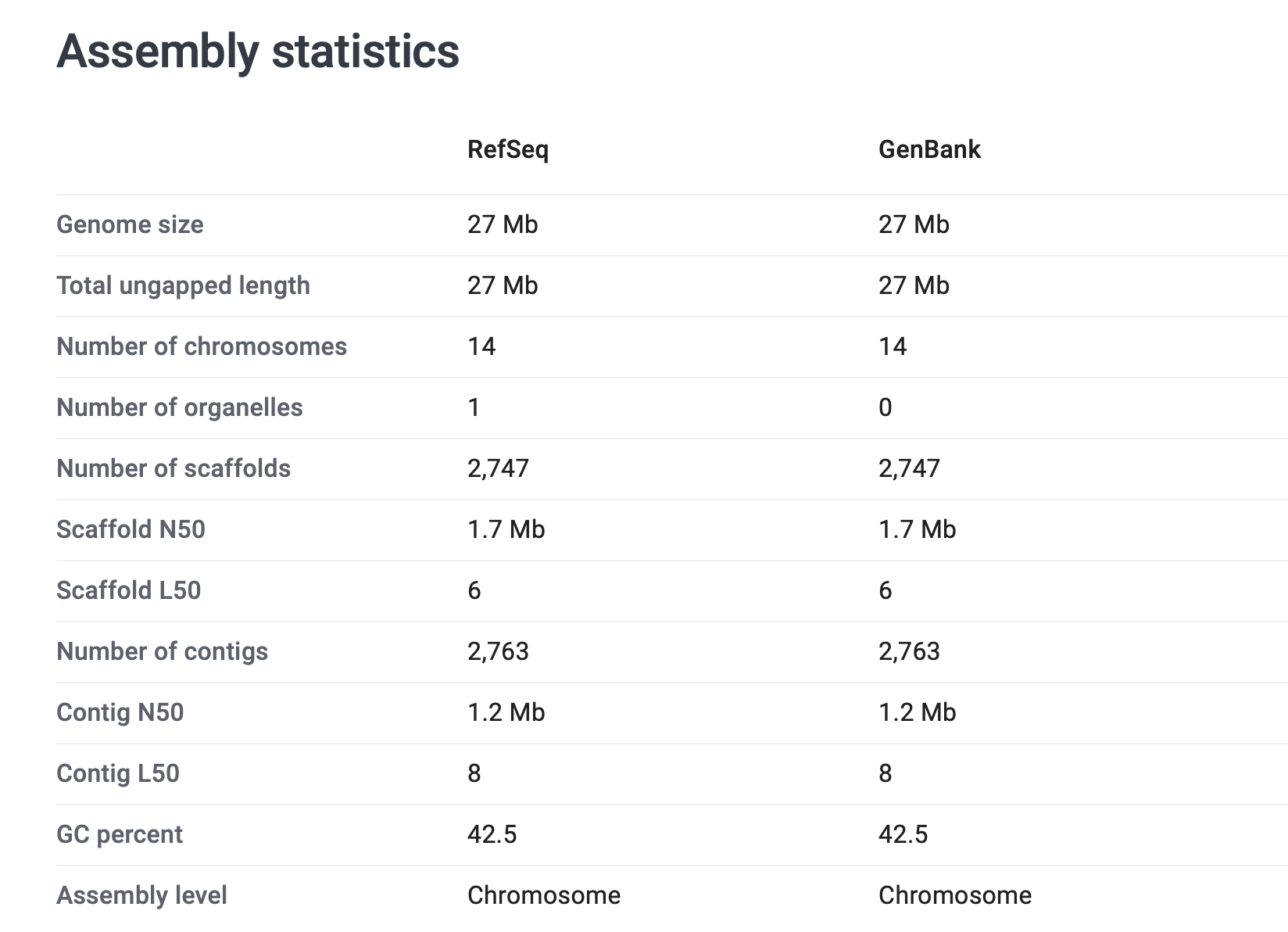
In short the total genome size is 27Mb (check!) but some of this is in 2,747 'scaffolds' that are not chromosomes. (They appear in this GFF file as 'supercontig'.) These are, I think, bits of genome that have been assembled but the assemblers don't quite know where they should go. In principle they might be parts of nuclear chromosomes, parts of the mitochondria or other organelles like the apicoplast, or other bits of DNA that were sequenced but not clear where they should go.
However other genomes, like Bufo bufo, seem to have only 'region' records even though several of those do seem to represent chromosomes. It seems that files from the Darwin Tree of Life project, like the Dromedary Camel genome, use 'region' entries. (That number 37 is not wrong - camels have 74 diploid chromosomes).
In any case - for our purposes, let's use the 'regions', 'chromosomes', and 'supercontigs' to represent the genome length.
Ok! Armed with chromosome lengths, we are ready to ask how much of the genome is in genes?