What's not a gene?
Above we counted all "genes". But what exactly is a "gene"?
One way to figure out what 'gene' means in these files, is to compare to the other record types in the data. For
example, you can find all the top-level record types (those with no Parent) in the file like this:
- sqlite3 code
- R code
- dbplyr code
- Python code
sqlite> SELECT dataset, type, COUNT(*) FROM gff WHERE Parent IS NULL GROUP BY dataset, type
(
dbGetQuery( db, "SELECT * FROM gff WHERE Parent IS NULL" )
%>% group_by( dataset, type )
%>% summarise( count = n() )
)
(
db
%>% tbl( 'gff' )
%>% filter( is.na(Parent ))
%>% group_by( dataset, type )
%>% summarise( count = n() )
%>% collect()
)
types = pandas.read_sql(
"SELECT dataset, type, COUNT(*) FROM gff WHERE Parent IS NULL GROUP BY dataset, type",
db
)
Note
For the example above, and those from now on, we will assume you have already connected to the database as we did on the previous page. Depending on how you are running this, you can connect like this:
- sqlite3 code
- R code
- Python code
sqlite3 genes.sqlite
sqlite3> .mode column
sqlite3> .header on
library( RSQLite )
library( dplyr )
db = DBI::dbConnect( RSQLite::SQLite(), "genes.sqlite" )
import pandas, sqlite3
db = sqlite3.connect( "genes.sqlite" )
If not, please do that now and then re-run the above.
Whichever way you do this, you should see that all the files have a bunch of records other than protein-coding genes. There are, at least, pseudogenes and non-coding RNAs, as well as a bunch of so-called biological regions and untranslated regions.
Question
What are pseudogenes and non-coding RNAs anyway? To see the technical definitions, look them up on the sequence ontology.
For more information, here is an interesting reference on pseudogenes and on non-coding RNAs.
Different types of gene
What sets apart genes from pseudogenes and non-coding RNAs is that they code for proteins. However, it's not quite
that simple. If you look more closely at the records with type=gene, you'll see they have a number of different
"biotypes" (which, if you've followed so far, you will have avialable in the biotype column.)
Challenge
Extend the query above to count records by both type and biotype.
What types of gene record are there? What types of non-coding RNA? What types of pseudogene?
The sequence ontology page is a good first place to look these up if you want to know more about them.
Aside on T and B cell receptors
One of the record types you'll see looks like genes called 'IG_C_gene', 'IG_D_gene', and so on. And some are called 'TR_C_gene' and so on.
These are immunoglobulin and T cell receptor gene segments respectively. The mechanism that turns these gene segments into functional, expressed genes that code for proteins - known as V(D)J recombination - is complex, and involves somatic structural variation that brings 'V_gene', 'J_gene', and at some loci 'D_gene' segments together to form fully functional genes. This only happens in B cells and T cells and is a basic component of the adaptive immune system.
For example, here is a cartoon of the T cell receptor 'alpha' and 'delta' regions from the T cell Receptor Factsbook:
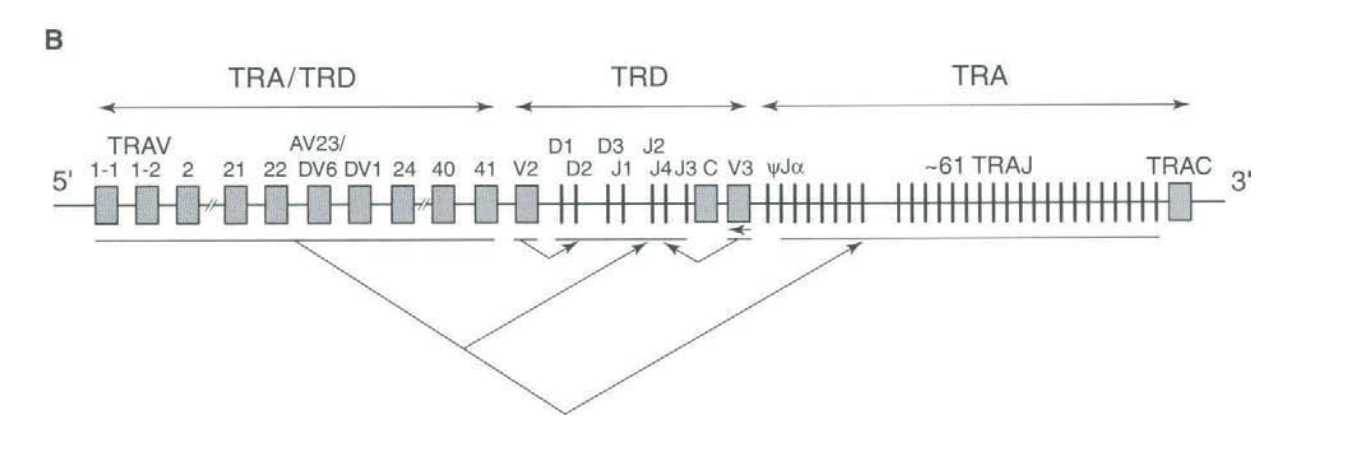
The arrows underneath indicate a schema of how the 'V' and 'J' segments are joined together, or the 'V', 'D', and 'J' segments for the TRD genes. The component genes are chosen randomly in each cell to generate a high level of receptor diversity.
Note
If that's not complicated enough - the diagram also shows that the TRD locus is situated inside the TRA locus. Sometimes they also recombine together to make hybrid A/D genes.
In any case - although these genes do code for proteins, they aren't standard 'protein coding genes' in that they don't form full genes in most cells - only after recombination in T and B cells. For that reason they are seperated out from protain coding genes in the data.